GITT analysis#
In this notebook we will use cellpy to extract the open circuit voltages (OCV) from a GITT measurement. The extracted OCVs will be plotted, and the results saved in .csv format.”
[1]:
import pathlib
import pandas as pd
import matplotlib.pyplot as plt
import cellpy
from cellpy.utils import plotutils
Set filepath and load the datafile:
[2]:
filedir=pathlib.Path("data") # foldername within the same directory
c = cellpy.get(filedir / 'out' / '20210210_FC.h5')
Produce an overview plot to identify cycle numbers for the GITT experiment (for an interactive version of this plot, you have to have plotly installed):
[3]:
plotutils.cycle_info_plot(c, cycle = list(range(2, 7)))
OCV extraction#
From the overview plot above, we can identify the GITT cycles to be cycle number 4 and 5. In the following, we will focus on cycle 5 only.
For further analysis, we create the step table, called steps, a dataframe that contains a lot ofnformation on all the cycle steps for the cell.
In the following, we apply several filters to steps, to eventually extract OCV voltages and corresponding capacities:
``steps_cycle``: Extract the rows specifically for the selected GITT cycle (here: cycle Nr 5).
NB: For simplicity, steps_cycle only contains rows relevant for further analysis, i.e. “cycle”, “step””charge_last”, “discharge_last”, “voltage_first” ,”voltage_last”, “type”.”
[4]:
GITT_cycle=5
c.make_step_table(all_steps=True)
steps=c.data.steps
steps_cycle=steps.loc[(steps.cycle==GITT_cycle),["cycle", "step", "charge_last", "discharge_last", "voltage_first" ,"voltage_last", "type"]]
Taking a closer look at the created steps_cycle dataframe:
steps_cycle.head(10)to view the first 10 rowssteps_cycle.tail(10)to view the last 10 rows
[5]:
steps_cycle.tail(10)
[5]:
| cycle | step | charge_last | discharge_last | voltage_first | voltage_last | type | |
|---|---|---|---|---|---|---|---|
| 755 | 5 | 8 | 0.003358 | 0.003258 | 3.212396 | 3.343531 | ocvrlx_up |
| 756 | 5 | 7 | 0.003358 | 0.003294 | 3.330632 | 3.139919 | discharge |
| 757 | 5 | 8 | 0.003358 | 0.003294 | 3.162645 | 3.314970 | ocvrlx_up |
| 758 | 5 | 7 | 0.003358 | 0.003330 | 3.302993 | 3.080647 | discharge |
| 759 | 5 | 8 | 0.003358 | 0.003330 | 3.102759 | 3.283338 | ocvrlx_up |
| 760 | 5 | 7 | 0.003358 | 0.003366 | 3.272282 | 3.008170 | discharge |
| 761 | 5 | 8 | 0.003358 | 0.003366 | 3.029361 | 3.246485 | ocvrlx_up |
| 762 | 5 | 7 | 0.003358 | 0.003392 | 3.233587 | 2.999878 | discharge |
| 763 | 5 | 10 | 0.003358 | 0.003392 | 3.010627 | 3.010627 | ir |
| 764 | 5 | 11 | 0.003358 | 0.003392 | 3.037038 | 3.228980 | ocvrlx_up |
To extract the OCV voltages, we then filter the
steps_cycledataframe forthe OCV relaxation steps on discharge,
steps_ocv_dch, of type oxvrlx_up (and rest), corresponding tostep==3, andthe OCV relaxation steps on charge
steps_ocv_cha, of type oxvrlx_down (and rest), corresponding tostep==8. Thereby we obtain two new dataframes
[6]:
steps_ocv_cha = steps_cycle.loc[steps_cycle.step==3]
steps_ocv_dch = steps_cycle.loc[steps_cycle.step==8]
[7]:
steps_ocv_cha.head(5)
[7]:
| cycle | step | charge_last | discharge_last | voltage_first | voltage_last | type | |
|---|---|---|---|---|---|---|---|
| 390 | 5 | 3 | 0.000036 | 0.0 | 3.512440 | 3.487564 | rest |
| 392 | 5 | 3 | 0.000072 | 0.0 | 3.518582 | 3.494320 | rest |
| 394 | 5 | 3 | 0.000109 | 0.0 | 3.524724 | 3.499848 | rest |
| 396 | 5 | 3 | 0.000145 | 0.0 | 3.530559 | 3.505991 | rest |
| 398 | 5 | 3 | 0.000181 | 0.0 | 3.537315 | 3.513054 | rest |
The voltages at the end of these steps (voltage_last), contain the (pseudo-) OCV voltages:
[8]:
V_cha = steps_ocv_cha.voltage_last.reset_index(drop=True)
V_dch = steps_ocv_dch.voltage_last.reset_index(drop=True)
cap_cha= steps_ocv_cha.charge_last.reset_index(drop=True)*1000 #*1000 to convert to mAh
cap_dch= steps_ocv_dch.discharge_last.reset_index(drop=True)*1000 #*1000 to convert to mAh
To plot our results, we additionally get the entire voltage vs capacity curves for the selected GITT cycle, employing the .get_ccap and .get_dcap methods. The cell mass is used to convert from gravimetric capacity (mAh/g) to capacity (mAh).
[9]:
c.make_step_table(all_steps=False)
ccap = c.get_ccap(cycle=GITT_cycle)
dcap = c.get_dcap(cycle=GITT_cycle)
mass = c.get_mass() # in mg
[10]:
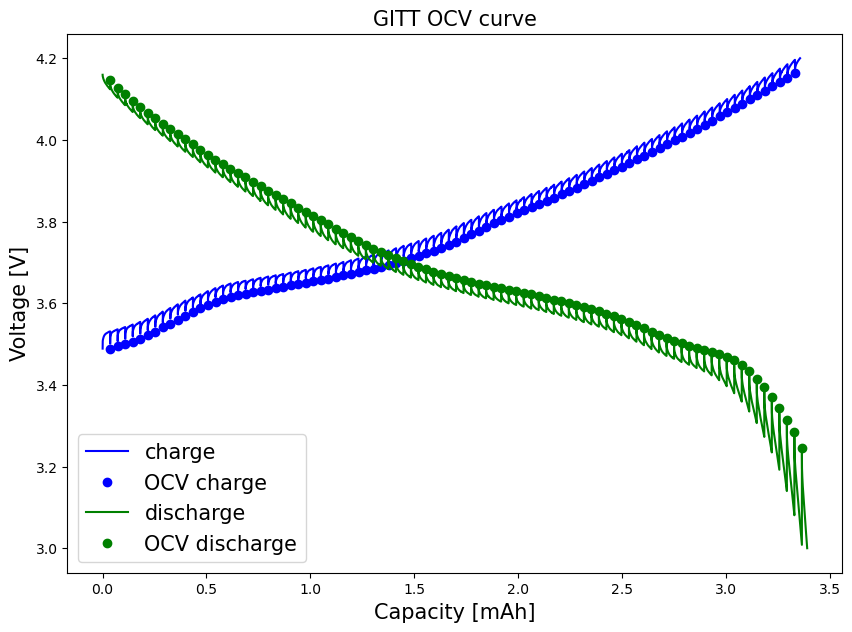
fig, ax = plt.subplots()
ax.plot(ccap["charge_capacity"]*mass/1000, ccap["voltage"], color='blue', label="charge")
ax.plot(cap_cha, V_cha, "bo", label="OCV charge")
ax.plot(dcap["discharge_capacity"]*mass/1000, dcap["voltage"], color="green", label="discharge")
ax.plot(cap_dch, V_dch, "go", label="OCV discharge")
plt.xlabel("Capacity [mAh]", fontsize=15)
plt.ylabel("Voltage [V]", fontsize=15)
plt.title("GITT OCV curve", fontsize=15)
#plt.ylim(0, 0.91)
#plt.xlim(0, 4.70)
ax.legend(fontsize=15)
fig.set_figheight(7)
fig.set_figwidth(10)
plt.show()
Saving the data#
Concatenate the OCV voltages and capacities into a dataframe, and save as a .csv file.
[11]:
OCV_cha = pd.concat([cap_cha, V_cha], axis=1 ,keys=["Charge_cap_mAh","OCV_V"])
OCV_dch = pd.concat([cap_dch, V_dch], axis=1, keys=["Discharge_cap_mAh", "OCV_V"])
[12]:
#OCV_cha.to_csv('GITT_OCV_cycle'+str(GITT_cycle)+'_cha.csv', index=False)
#OCV_dch.to_csv('GITT_OCV_cycle'+str(GITT_cycle)+'_dch.csv', index=False)